# Libraries
library(tidyverse)
library(tidymodels)
# Ensuring tidymodels functions priority
tidymodels_prefer()
# Importing customer_churn
customer_churn <- read_csv("https://raw.githubusercontent.com/Datakortex/Datasets/refs/heads/main/customer_churn.csv")
# Importing and pre-processing the dataset
customer_churn <- customer_churn %>%
select(Recency, Frequency, Monetary_Value, Churn) %>%
mutate(Churn_Label = as.factor(if_else(Churn == 1, "Churn", "No Churn")))
# Setting seed
set.seed(999)
# Splitting customer_churn
split_data <- initial_split(customer_churn, prop = 0.75, strata = Churn_Label)
# Extracting the training set
training_set <- split_data %>% training()
# Extracting the test set
test_set <- split_data %>% testing()
# 5x3 (repeated) cross-validation folds
folds_rep_cv <- vfold_cv(training_set, v = 5, repeats = 3)Hyperparameter Tuning
Introduction
In Chapter Introduction to Machine Learning, we mentioned that most machine learning models come with hyperparameters, which are settings that control the learning process. In fact, nearly all machine learning models include some form of hyperparameter. Even the Laplace estimator in a Naive Bayes model can be interpreted as a hyperparameter (see Chapter Naive Bayes).
In this chapter, we focus on one of the simplest yet most effective approaches for selecting these values in order to improve predictive performance. We discuss how to tune hyperparameters using tidymodels, how this process is integrated into workflows, and how different hyperparameter settings can be compared in terms of predictive performance.
Hyperparameter Tuning with Regular Grid Search
The goal of hyperparameter tuning is to improve model performance on unseen data. In previous chapters, we introduced the main hyperparameters for each machine learning method, but we typically assigned them fixed values without extensive justification. This was partly for simplicity, but also reflects the idea behind the No Free Lunch theorem: there is no universally optimal choice of hyperparameters. Because of this, hyperparameter tuning is an essential part of the modeling workflow. It allows us to adapt models to the specific structure of the data at hand rather than relying on default or arbitrary choices.
There are different approaches to hyperparameter tuning, meaning methods for finding appropriate values that lead to good predictive performance. In this chapter, we focus on the simplest and most widely used approach: the regular grid search. The idea is to pre-specify a set of candidate values for each hyperparameter and evaluate all possible combinations using resampling methods. Each combination is treated as a separate model, and its performance is estimated using, for example, cross-validation. The final choice is then based on the average performance across resamples.
For instance, in the case of a k-nearest neighbors (KNN) model, the main hyperparameters is the number of neighbors. A common rule of thumb is to set \(k\) to the square root of the number of observations. However, instead of relying on a single heuristic value, grid search allows us to systematically evaluate a range of values around this guideline. For example, if we have a dataset with 3000 observations, the square-root rule suggests a value of approximately 55. Using a regular grid approach, we would not only test this value, but also a set of nearby values, such as 10, 25, 50, 75, 100, and 150, to assess how sensitive model performance is to the choice of k.
The main advantage of this approach is its simplicity. It is easy to understand and implement, and integrates directly with the resampling framework introduced in the previous chapter. However, the main drawback is computational cost. As the number of hyperparameters increases, the total number of combinations grows multiplicatively. This can quickly become infeasible, especially for complex models or large datasets. For example, if we tune three hyperparameters and consider only five values for each, we already end up with 125 possible configurations, each of which would need to be evaluated using cross-validation. For this reason, grid search is often used with relatively small and carefully chosen parameter ranges rather than exhaustive exploration.
From a practical perspective, it is suggested to start with a relatively wide grid that covers a reasonable range of values for each hyperparameter. At this stage, the goal is not to find the exact optimum, but to understand how performance behaves across different regions of the parameter space. Once we identify values that lead to better performance, we can refine the grid by focusing on a narrower range with finer resolution.
As with resampling methods, it is important that hyperparameter tuning is performed entirely only within the training data. The test set should not be used during this process, as this would again introduce optimistic bias. Instead, we rely on validation set(s) or cross-validation performance to guide the selection of hyperparameters. Once the final configuration has been selected, the model is refit on the full training data using the chosen hyperparameters. This ensures that the model incorporates all available information before being evaluated on the test set, which remains the final and unbiased measure of performance.
Hyperparameter Tuning with Tidymodels
With tidymodels, hyperparameter tuning is implemented using workflows. To see how this works, we load the tidyverse and tidymodels packages and import the customer_churn dataset, as in Chapter Introduction to Tidy Modeling. We then use initial_split(), training(), and testing() to create training and test sets, and vfold_cv() to generate 5×3 repeated cross-validation folds:
We now define a KNN model. Instead of setting the number of neighbors directly, we use tune() to indicate that this value will be optimized later. At this stage, nothing is fitted yet. We are only defining the model structure, and leaving the hyperparameter open for tuning. We also define a recipe to normalize the predictors and combine everything into a workflow:
# Initiating engine using tune()
knn_spec <- nearest_neighbor(neighbors = tune()) %>%
set_engine("kknn") %>%
set_mode("classification")
# Creating recipe
knn_recipe <- recipe(Churn_Label ~ Recency + Frequency + Monetary_Value,
data = training_set) %>%
step_normalize(all_numeric_predictors())
# Combining model and recipe into a workflow
knn_workflow <- workflow() %>%
add_model(knn_spec) %>%
add_recipe(knn_recipe)
# Printing workflow output
knn_workflow══ Workflow ═════════════════════════════════════════════════════
Preprocessor: Recipe
Model: nearest_neighbor()
── Preprocessor ─────────────────────────────────────────────────
1 Recipe Step
• step_normalize()
── Model ────────────────────────────────────────────────────────
K-Nearest Neighbor Model Specification (classification)
Main Arguments:
neighbors = tune()
Computational engine: kknn The output shows that neighbors is now set to tune(), meaning the value will be chosen during the tuning step.
Since we are tuning this parameter, we need to define a set of candidate values. A simple way is to create a tibble where the column name matches the hyperparameter name. We use the seq() function to generate a sequence of values, which is convenient because it avoids manually specifying each value. In our case, we generate values from 5 to 105 in steps of 10:
# Creating tibble with the chosen hyperparameter values
knn_grid <- tibble(
neighbors = seq(from = 5, to = 105, by = 10)
)We now apply cross-validation using tune_grid(). This replaces fit_resamples() from earlier chapters. The difference is that now we evaluate multiple hyperparameter values instead of fitting a single fixed model. To do this, we also include the additional argument grid, where we pass the tibble we created. This tells the function which hyperparameter values should be evaluated during resampling.
# Setting seed
set.seed(123)
# Fitting resamples and hyper
knn_results <- knn_workflow %>%
tune_grid(
resamples = folds_rep_cv,
grid = knn_grid,
metrics = metric_set(precision, recall, f_meas))
# Printing knn_results
collect_metrics(knn_results)# A tibble: 33 × 7
neighbors .metric .estimator mean n std_err .config
<dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
1 5 f_meas binary 0.679 15 0.00738 pre0_mod01…
2 5 precision binary 0.720 15 0.0108 pre0_mod01…
3 5 recall binary 0.644 15 0.00992 pre0_mod01…
4 15 f_meas binary 0.707 15 0.00767 pre0_mod02…
5 15 precision binary 0.777 15 0.0109 pre0_mod02…
6 15 recall binary 0.650 15 0.0105 pre0_mod02…
7 25 f_meas binary 0.706 15 0.00714 pre0_mod03…
8 25 precision binary 0.783 15 0.0105 pre0_mod03…
9 25 recall binary 0.644 15 0.00852 pre0_mod03…
10 35 f_meas binary 0.706 15 0.00777 pre0_mod04…
# ℹ 23 more rowsWe can create plots to visualize the results using the function autoplot() on the resulting object. This function essentially uses ggplot2 under the hood and automatically produces a summary plot of the resampling results across the tested hyperparameter values:
# Plotting modeling results
autoplot(knn_results)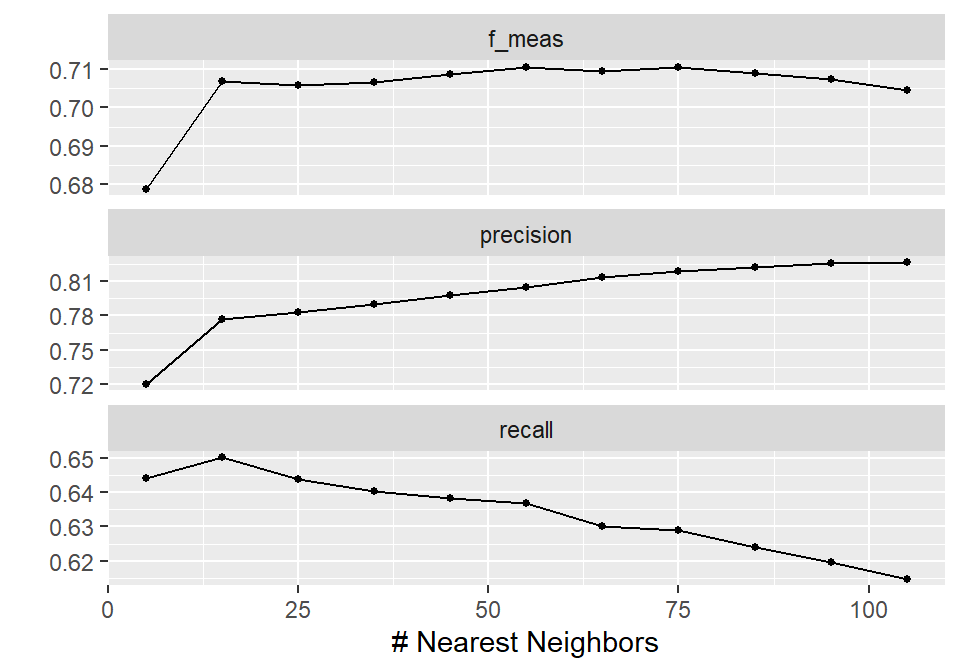
In this way, the results are shown across different values of neighbors, making it easy to compare model performance. Based on this visualization, 55 neighbors leads to the highest F1-score, with a precision slightly above 80% and a recall close to 64%. In practical terms, this means that around 80% of the predicted churners are actually correct, while about 64% of all true churners are identified by the model.
Note that these are average values across the resamples, but based on our modeling setup, 55 neighbors appears to be a reasonable choice if this trade-off is acceptable from a business perspective. Interestingly, this choice also aligns with the common rule of thumb of using the square root of the number of observations as the number of neighbors in a KNN model.
We can extract the best-performing hyperparameters using the function select_best(). In this function, we set the argument metric to the metric of interest. If we do not specify it, the first metric listed in metric_set() is used by default:
# Selecting best model
best_params <- knn_results %>%
select_best(metric = "f_meas")
# Printing best_params
best_params# A tibble: 1 × 2
neighbors .config
<dbl> <chr>
1 55 pre0_mod06_post0We can now finalize the workflow using the function finalize_workflow(). This step takes the workflow where the hyperparameter has been left unspecified using tune()—which acts as a placeholder rather than an actual value—and replaces it with the selected optimal value from the tuning results. In other words, we move from a model definition where a position is reserved for a hyperparameter to a fully specified model where all values are fixed and ready for fitting as the final model. Note that no model fitting takes place at this stage; it only prepares the workflow so that it can be trained with the chosen hyperparameter setting:
# Finalizing the workflow
final_workflow <- finalize_workflow(x = knn_workflow, parameters = best_params)To fit the final model on the full training set and evaluate it on the test set, we could use the fit() function as in previous chapters. However, there is also a more convenient approach in this workflow setting using the function last_fit(). This function automatically refits the finalized workflow on the training data and evaluates it on the test set in a single step. We just pass the final_workflow object and the split_data object we created earlier. Once again, we specify the metrics argument to define how model performance is evaluated on the test set. We use the same metrics as before, and additionally include roc_auc to measure the model’s ability to distinguish between the two classes across all classification thresholds.
Disclaimer
It is strongly recommended to use the same set of metrics during both the training and testing phases to ensure consistency in model evaluation. In this case, we additionally include
roc_aucat the testing stage to illustrate how ROC-based evaluation can be computed and visualized within the tidymodels framework, even though it was not part of the tuning metrics.
# Fitting the model on the training set and evaluating it on the test set
final_fit <- last_fit(
object = final_workflow,
split = split_data,
metrics = metric_set(roc_auc, precision, recall, f_meas)
)
# Collecting metrics
collect_metrics(final_fit)# A tibble: 4 × 4
.metric .estimator .estimate .config
<chr> <chr> <dbl> <chr>
1 precision binary 0.843 pre0_mod0_post0
2 recall binary 0.688 pre0_mod0_post0
3 f_meas binary 0.758 pre0_mod0_post0
4 roc_auc binary 0.918 pre0_mod0_post0The test metrics are slightly lower than the cross-validation results, which is expected. Some drop in performance is normal, since cross-validation provides an estimate based on resamples, while the test set represents completely unseen data.
Lastly, we can use an ROC curve plot, a concept we discussed in Chapter Model Validation and Performance Evaluation. To do this, we first need to collect the predictions using the collect_predictions() function on the final_fit object. We then use the function roc_curve(), which computes the true positive and false positive rates across different probability thresholds and constructs the ROC curve. In this function, we specify the true class labels using the truth argument and the predicted probabilities for the positive class (e.g., .pred_Churn) as the input, which are used to calculate the curve. To finalize the plot, we use the autoplot() function:
# Collecting predictions
knn_predictions <- collect_predictions(final_fit)
# Plotting ROC curve
roc_curve(knn_predictions, truth = Churn_Label, .pred_Churn) %>%
autoplot()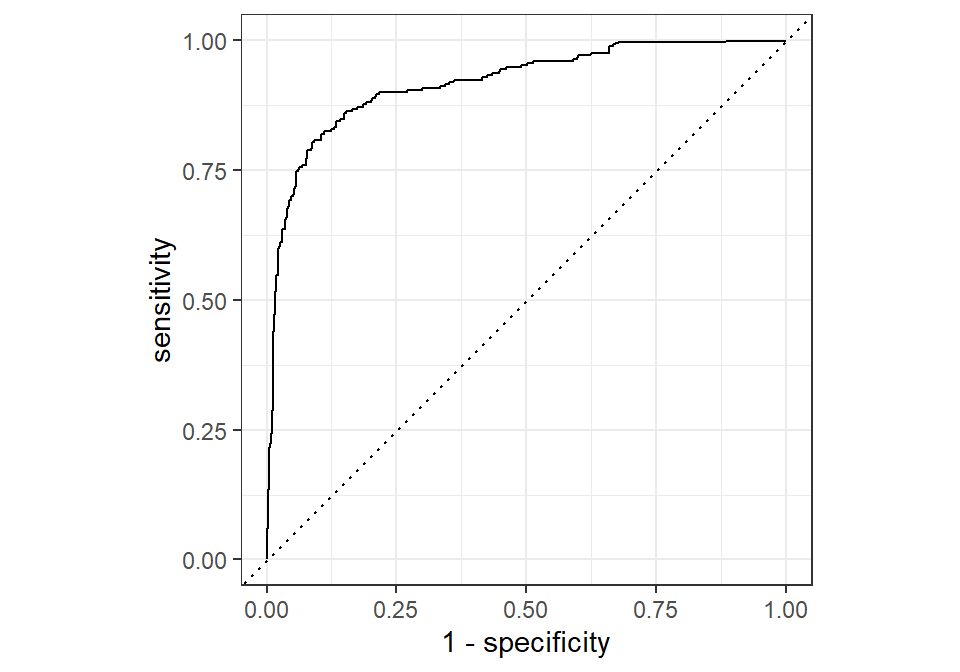
Recap
In this chapter, we focused on hyperparameter tuning using regular grid search. The idea is to systematically evaluate multiple combinations of hyperparameter values using resampling methods and select the configuration that performs best on average. We showed how this process is implemented in tidymodels using workflows, tune(), and tune_grid(), and how the tuning process is integrated with cross-validation in a consistent framework.
We also highlighted the trade-off between simplicity and computational cost. While grid search is easy to understand and implement, it can quickly become expensive as the number of hyperparameters and candidate values increases. Finally, we demonstrated how the best-performing model is selected, finalized, and evaluated on a held-out test set to obtain an unbiased estimate of performance.